Unwrap a blade’s profile¶
Description¶
This treatment unwraps a line’s profile.
It must be applied on a 1D line resulting from a cut of a 3D blade.
As a result, the indices of nodes from pressure and suction sides will be available as attributes of the base.
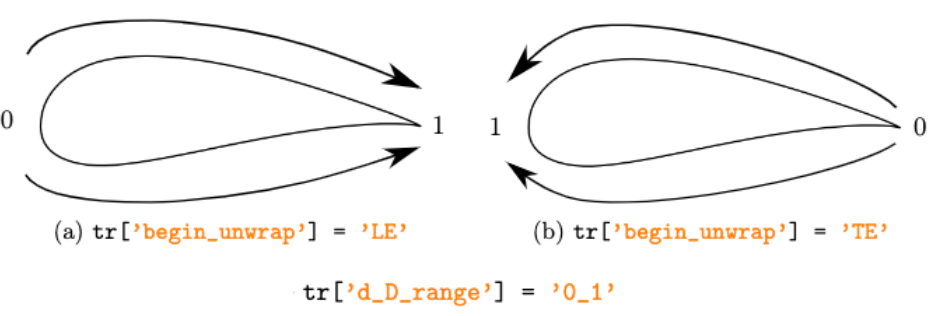
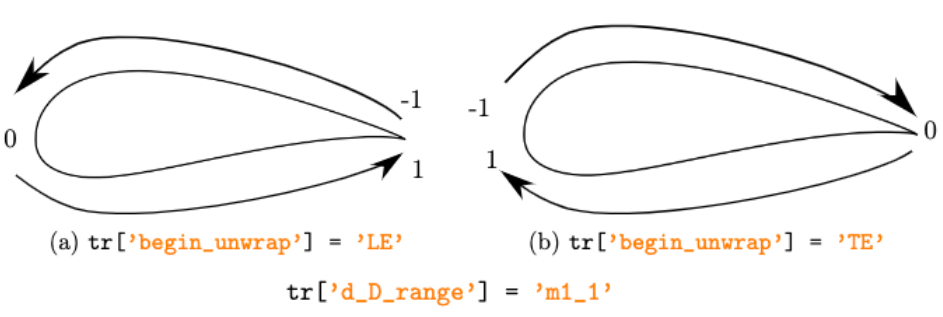
Examples of begin_unwrap and d_D_range options.
Parameters¶
- base:
Base The base on which the treatment will be applied.
- base:
- p_var:
str, default = ‘Psta’ Name of pressure variable.
- p_var:
- d_D_range:
strin [‘m1_1’, ‘0_1’], default = ‘0_1’ Range of the curvilinear reduced coordinates. Either ‘m1_1’ for a range from -1 to 1 or ‘0_1’ for a range from 0 to 1.
- d_D_range:
- begin_unwrap:
strin [‘LE’, ‘TE’], default = ‘LE’ Convention: ‘LE’ (leading edge) or ‘TE’ (trailing edge). Either sort the blade from the LE to the TE or vice versa. Therefore, if begin_unwrap is set to ‘LE’, the origin of the curvilinear coordinate is at the leading edge.
- begin_unwrap:
- tolerance_decimals:
int, default = None Number of decimal places to round the coordinates. When the base is multi-zone, it is used to merge the intersection points of the zones. If None, the coordinates will not be rounded.
- tolerance_decimals:
- memory_mode:
bool, default = False If True, the input base is deleted on the fly to limit memory usage.
- memory_mode:
- cartesian_coordinates:
list(str, str, str), default = [‘x’, ‘y’, ‘z’] Names of the 3 Cartesian coordinates.
- cartesian_coordinates:
- crm_var:
str, default = ‘CoordinateReducedMeridional’ Name of the reduced meridional coordinate variable.
- crm_var:
- out_vars:
sequence(str, str), default = [‘d_D’, ‘d’] Name of the output reduced curvilinear curvature coordinate and its non reduced form.
- out_vars:
Preconditions¶
The treatment must preferably be applied on a 1D Zone resulting from
a blade cut. However, a merge treatment can handle a base with multiple zones.
The Zone must contain only one Instant.
Postconditions¶
The input base may be modified in-place depending on the result of the inner merge treatment.
The output base is a merge of the input base extended with the attribute
named ‘Profil’ in Base.attrs.
This attribute is a dictionary with variables:
- IndicesPressureSide
Instant’s nodes indices corresponding to the profile’s pressure side.
- IndicesSuctionSide
Instant’s nodes indices corresponding to the profile’s suction side.
The Instant contained in the input base is extended with the following variables:
- out_vars[1]
The curvilinear curvature coordinate of the profile.
- out_vars[0]
Reduced form of the curvilinear curvature coordinate. In other words, the curvilinear curvature coordinate of the profile divided by the total curvilinear curvature length.
Example¶
import antares
myt = antares.Treatment('UnwrapProfil')
myt['base'] = base
base = myt.execute()
The following statements plot the temperature as a function of the reduced curvilinear abscissa coordinate.
import matplotlib.pyplot as plt
# First, retrieve pressure and suction sides indices:
ind_PS = base.attrs['Profil']['IndicesPressureSide']
ind_SS = base.attrs['Profil']['IndicesSuctionSide']
# Get variables you want to plot
T = base[0][0]['T']
d_D = base[0][0]['d_D']
# Create a basic figure
plt.figure()
plt.plot(d_D[ind_PS], T[ind_PS], label='Pressure side')
plt.plot(d_D[ind_SS], T[ind_SS], label='Suction side')
plt.legend()
plt.show()